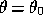 , then the likelihood function can be approximated by
another multivariate Gaussian about this point;
, then the likelihood function can be approximated by
another multivariate Gaussian about this point;
Suppose we have found the maximum likelihood solutions for each parameter,
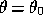 , then the likelihood function can be approximated by
another multivariate Gaussian about this point;
, then the likelihood function can be approximated by
another multivariate Gaussian about this point;
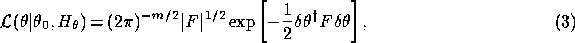
where 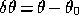 is the distance to the maximum in
parameter space and the parameter covariance matrix is given by the inverse of
is the distance to the maximum in
parameter space and the parameter covariance matrix is given by the inverse of
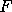 , the Fisher Information matrix;
, the Fisher Information matrix;

(if the means of the data are dependent on the parameters, this is modified --
see Tegmark, Taylor & Heavens (1997)). The far right hand side expression can be calculated for
Gaussian distributed data sets (ie equation 1), where 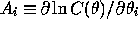 is the slope of the log of the data covariance
matrix in parameter space.
is the slope of the log of the data covariance
matrix in parameter space.
By considering the Fisher matrix as the information content contained in the data set about each parameter, we see that the solution to our problem is to reduce the data set without changing the parameter information content. Hence to solve the problem of efficiency, we need to make a linear transformation of the data set

where 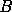 is a
is a 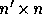 matrix where
matrix where 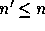 , and so
, and so 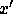 may be a
smaller data set than
may be a
smaller data set than  . If
. If 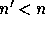 the transformation is not invertible
and some information about the data has been lost. To ensure that the lost
information does not affect the parameter estimation (requirement (a)), we also
require
the transformation is not invertible
and some information about the data has been lost. To ensure that the lost
information does not affect the parameter estimation (requirement (a)), we also
require

where 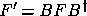 is the transformed Fisher matrix. In order to avoid
learning the unhelpful fact that no data is an optimal solution, we add in the
constraint that data exists. Since we have the freedom to transform the data
covariance matrix, we add the constraint
is the transformed Fisher matrix. In order to avoid
learning the unhelpful fact that no data is an optimal solution, we add in the
constraint that data exists. Since we have the freedom to transform the data
covariance matrix, we add the constraint 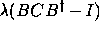 , where
, where 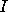 is
the unit matrix and
is
the unit matrix and 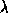 is a Lagrangian multiplier.
is a Lagrangian multiplier.
It can be shown (Tegmark, Taylor & Heavens (1997)) that this is equivalent to a generalised
Karhunen-Loève eigenvalue problem, which has a unique solution 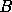 for each
parameter. These solutions have the property that
for each
parameter. These solutions have the property that

where 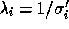 are the eigenvalues of the transformed data set
and the inverse errors associated with each eigenmode of the new data set.
are the eigenvalues of the transformed data set
and the inverse errors associated with each eigenmode of the new data set.
The new, compressed data set, 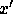 , can now be ordered by decreasing
eigenvalue, so that the first eigenmode contains the most information about the
desired parameter, the second slightly less information, and so on. The total
error on the parameter is then simply given by the inverse of the
, can now be ordered by decreasing
eigenvalue, so that the first eigenmode contains the most information about the
desired parameter, the second slightly less information, and so on. The total
error on the parameter is then simply given by the inverse of the 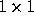 Fisher matrix
Fisher matrix

We are now free to choose how many eigenmodes to include in the likelihood
analysis. A compression of 10 will lead to a time saving of 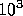 . However
this is only exact if we know the true value of the parameters used to
calculate
. However
this is only exact if we know the true value of the parameters used to
calculate 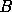 . But if we are near the maximum likelihood solution then we can
iterate towards the exact solution.
. But if we are near the maximum likelihood solution then we can
iterate towards the exact solution.
This procedure is optimal for all parameters -- linear and nonlinear -- in the
model. In the special case of linear parameters that are just proportional to
the signal part of the data covariance matrix (for example the amplitude of
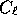 , if the data are the
, if the data are the 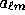 ), the eigenmodes reduce to
signal-to-noise eigenmodes (Bond (1994)). Hence our eigenmodes are more general
than signal-to-noise eigenmodes. Furthermore, as our eigenmodes satisfy the
condition that the Fisher matrix is a maximum, they are the optimal ones for
data compression. Any other choice, including signal-to-noise eigenmodes, would
give a higher variance.
), the eigenmodes reduce to
signal-to-noise eigenmodes (Bond (1994)). Hence our eigenmodes are more general
than signal-to-noise eigenmodes. Furthermore, as our eigenmodes satisfy the
condition that the Fisher matrix is a maximum, they are the optimal ones for
data compression. Any other choice, including signal-to-noise eigenmodes, would
give a higher variance.
In Figure 1 we plot the uncertainty on 3 parameters for COBE-type data,
the quadrupole, 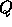 , the spectral index of scalar perturbations,
, the spectral index of scalar perturbations,  and the
re-ionization optical depth,
and the
re-ionization optical depth,  .
.
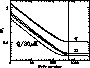
Figure 1: The 3 heavy lines show the error bars on 3 CMB
parameters as a function of the number of modes used. Each set of modes has
been optimised for the parameter in question. Note that approximately 400
modes are all that is required to get virtually all the information from the
entire 4016 cut COBE dataset. The thin lines show the conditional errors from
the SVD procedure outlined in section 5: virtually all the (conditional)
information on all 3 parameters is obtained from the best 500 SVD modes.